
Building a COVID-19 vulnerability index
Introduction
Coronaviruses (CoV) are a large family of viruses that cause illnesses ranging from the common cold to more severe diseases such as Middle East respiratory syndrome (MERS-CoV) and severe acute respiratory syndrome (SARS-CoV). CoV are zoonotic, meaning they are transmitted between animals and people. Coronavirus disease 2019 (COVID-19) is caused by a new strain discovered in 2019, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), that has not been previously identified in humans (1).
COVID-19 is a highly contagious respiratory infection with common signs that include respiratory symptoms, fever, cough, shortness of breath, and breathing difficulties. In more severe cases, infection can cause pneumonia, severe acute respiratory syndrome, kidney failure, cardiac arrest, and death (2,3). Experts continue to learn more about COVID-19, including its etiology, symptoms, complications, and potential treatments.
On March 11, 2020, the World Health Organization (WHO) declared COVID-19 to be a pandemic (4). Public health and healthcare experts agree that mitigation is required in order to slow the spread of COVID-19 and prevent the collapse of healthcare systems. Health systems in the US run close to capacity, and so every transmission that can be avoided and every case that can be prevented has enormous impact (5).
The risk of severe complications from COVID-19 is higher for certain vulnerable populations, particularly people who are elderly, frail, or have multiple chronic conditions. The risk of death has been difficult to calculate, but a small study of people who contracted COVID-19 in Wuhan along with patterns also seen in early reports from the US suggest that the risk of death increases with age, and is also higher for those who have diabetes, heart disease, blood clotting problems, or have shown signs of sepsis (6,7). With an average death rate of 1%, the death rate rose to 6% for people with cancer, high blood pressure, or chronic respiratory disease, 7% for people with diabetes, and 10% for people with heart disease (8). There was also a steep age gradient; the death rate among people aged 80 and over was 15% (9).
Identifying who is most vulnerable is not necessarily straightforward. More than 55% of Medicare beneficiaries meet at least one of the risk criteria listed by the US Centers for Disease Control and Prevention (CDC) (10). People with the same chronic condition don’t have the same risk, and many people will have other comorbidities, which compounds their vulnerability. Simple rules can’t reflect these differences or capture complex factors like frailty which makes people more vulnerable to severe infections (11).
Methods
Proxy outcome
Since real-world data on COVID-19 cases are not readily available, the C-19 index was developed using close proxy events. A person’s C-19 index is measured in terms of their near-term risk of severe complications from respiratory infections (e.g., pneumonia, influenza). The most direct proxy event is acute respiratory distress syndrome (ARDS), identified by the International Classification of Diseases version 10 (ICD-10) diagnosis code J80. ARDS is extremely rare, with an annual occurrence of less than 0.05% in Medicare members. To create a viable machine learning model, the outcome was broadened to include 4 closely related categories of respiratory diagnoses from the Clinical Classifications Software Refined (CCSR) classification system (12). Patients were considered to have the proxy event if they had any of the following diagnosis codes in any position on a medical claim associated with a hospital inpatient visit or observation stay:
- ICD-10-CM J80—ARDS;
- RSP002—pneumonia (except that caused by tuberculosis);
- RSP003—influenza;
- RSP005—acute bronchitis;
- RSP006—other specified upper respiratory infections.
Machine learning models were created that predict the likelihood that a patient will have an inpatient hospital stay due to one of the above conditions in the next 3 months. While claims-based machine learning models typically focus on longer term predictions such as outcomes within the next 6 months or 1 year, this model is expected to be used for immediate targeting decisions around COVID-19, and so a shorter window was deemed appropriate.
Data sets
These models were trained and initially tested using historical medical claims data. Two different data sets were obtained, representing different segments of the population. The first was the Center for Medicare & Medicaid Services (CMS) Limited Data Set (LDS) for 2015 & 2016 (13). The LDS contains beneficiary level health information for 5% of the Medicare population, and is available to the public subject to a Data Use Agreement. The second was medical claims data for 2.5 million beneficiaries obtained from Healthfirst, which provides health insurance for New Yorkers. These two data sets represent different demographic groups within the US population. The LDS data contains only Medicare beneficiaries, who are predominantly over the age of 65 or disabled. The Healthfirst data set is primarily a Medicaid population, which includes younger and more healthy adults.
In order to build a predictive model appropriate for the overall US population, cohorts were individually created for each data set and then the resulting cohorts were combined. The individual cohorts were designed so that when combined, they had an age profile consistent with the overall US population.
CMS data preparation
The CMS data cohort was created by identifying all living members above the age of 65 on 9/30/2016. This particular date was chosen because it was 3 months from the end of the data set and was therefore the latest possible prediction date in the data set. Using the latest date minimized the use of ICD-9 data in the prediction histories since the CMS data spanned the transition from ICD-9 to ICD-10 on October 1, 2015. Members in the CMS data cohort under 65 were excluded because the second population created using Healthfirst data was a better representation of adults under the age of 65. Only fee-for-service members were included because medical claims histories for other members are not reliably complete. We then excluded all members who had less than 6 months of continuous eligibility prior to 9/30/2016. We also excluded members who lost coverage within 3 months after 9/30/2016, except for those members who lost coverage due to death. These members are lost to follow-up. Since they lost eligibility, we would not see the proxy outcome in the data even if it occurred and exclude those members. Table 1 summarizes the population selection.
Table 1
| Population size | Selection criteria |
|---|---|
| 3,114,713 | Total members in the CMS dataset |
| 1,867,879 | Fee-for-service members with 6 months of continuous coverage prior to 9/30/2016 |
| 1,511,950 | 65 years old or older |
| 1,506,659 | Exclude members who died before 9/30/2016 |
| 1,500,700 | Exclude members who lose coverage before 12/31/2016 not due to death |
The selection criteria for the CMS population. The various inclusion criteria are listed along with the total number of members left after applying each criteria. CMS, Center for Medicare & Medicaid Services.
Healthfirst data preparation
The Healthfirst cohort contained a longer history, from 2017 to the present. For this data set members were evaluated at multiple points in time rather instead of having a single prediction date. Each person was evaluated using each month of eligibility as a prediction date. This approach was not used with the CMS data because that data didn’t cover a long enough time period to allow for multiple months. Members had to have 3 months of eligibility after and had to be at least 18 years old on the prediction date. Table 2 summarizes the population selection.
Table 2
| Population size | Selection criteria |
|---|---|
| 3,008,781 | Total members in the Healthfirst dataset |
| 30,898,086 | Total member-months of eligibility |
| 29,388,003 | 3 months of eligibility after the prediction date |
| 19,470,511 | 18 years or older on the prediction date |
The selection criteria for the Healthfirst population. The various inclusion criteria are listed along with the total number of members left after applying each criteria.
Combined population
Each data set was split by person 80%/20% into train and test sets. In the Healthfirst data set, doing the split by person ensures that all prediction dates for a given individual were in the same set, eliminating the possibility of data leakage. The data set for training was formed by taking all of the positive examples from each data set along with a sampling of the negative examples in the training set. The sampling was designed to meet two criteria. First, the percentage of the adult population ages 65 or older was set to match that of the US population (21% of adults). Second, the difference in the prevalence of the proxy outcome between those under and over 65 from the Healthfirst data set was maintained in the overall data set. This difference was 3.9×.
Despite these efforts, it is worth noting the ways in which this dataset differs from the general population of the US. Healthfirst is primarily a Medicaid population. The Medicaid population is demographically skewed toward the rules associated with Medicaid eligibility. Specifically, Medicaid has more representation from economically challenged, female, and disabled individuals than the general population. Healthfirst is also a geographically limited dataset, only from the New York area. This results in a population that is more urban, racially and ethnically diverse, and subject to a less diverse array of climates than the general population of the US. These differences are worth noting, given that these factors all have measurable correlations with health outcomes.
The combined test population was created by unioning the full test set from the CMS data and a random 20% sample of the Healthfirst test set. The full test population contained 1,621,149 training examples with a prevalence of 3.86%. The test population had 761,898 examples with a prevalence of 0.36%. The difference in prevalence is because negative examples were downsampled by 90% in the training set to reduce class imbalance. The testing set was not downsampled in order to provide accurate estimates of model performance on a realistic population.
Variable encoding
The models aim to leverage sources of information that are consistently available for those who have medical billing data. Patient age, count of emergency room (ER) visits, and count of inpatient admissions are features that are naturally numerically encoded. Gender is encoded using females as the base class (0= female, 1= male).
The most significant data preparation action is converting the patient’s diagnosis history into categorical data. This entails looking at all diagnosis codes occuring in the year leading up to the date of prediction and excluding the most recent 3 months due to the standard delays in processing claims data. From there, each diagnosis code is mapped to its corresponding CCSR categories. Boolean variables are created for each category indicating if the patient has at least one diagnosis from that category in their history.
Models
Our primary motivation in this effort was to equip healthcare professionals with an analytic tool to assess their population susceptibility to acute COVID-19 infections. In order to encourage wide adoption and rapid implementation of the predictive models, we created three separate models which represent different tradeoffs between accuracy and ease of implementation. All three models were trained and tested on the same data set. The models were built using the ClosedLoop platform on Amazon Web Services (AWS). The models are summarized in Table 3 below.
Table 3
| Model name | Algorithm | Deployment | Features | Sensitivity 5% (%) |
|---|---|---|---|---|
| Survey risk factors | Logistic regression | Online survey | 14 | 49.8 |
| Diagnosis history | XGBoost | Python package | 559 | 53.8 |
| Expanded feature | XGBoost | Hosted service | 892 | 54.1 |
A comparison of the models based on both deployment complexity and accuracy. For each model, the algorithm used, deployment mechanism, and total number of features are listed. Model performance is shown using the sensitivity at 5% alert rate, which indicates the percentage of vulnerable patients who are identified in the top 5% of predictions. The survey risk factors model has the simplest deployment mechanism and smallest number of features, but lower sensitivity than the more difficult to implement models.
The “survey risk factors” model is the simplest and uses only a small number of features that were designed to be able to be generated from a simple health risk assessment questionnaire. This model requires no technical implementation, and we have made it available through a web-based survey (14). The “diagnosis history” model uses 559 features derived from medical claims diagnosis and utilization data. We have made this model available on GitHub (15). Finally, the “expanded feature” model uses an extensive feature set derived from medical claims data along with linked geographical and social determinants of health data (16). This model is being made freely available to healthcare organizations. Information about accessing the platform can be found at https://cv19index.com.
Survey risk factors model
The first model is aimed at using a simple health history survey to enrich the high-level recommendations from the CDC website for identifying those individuals who are at risk (10). The CDC identifies risk factors as:
- Older adults;
- Individuals with heart disease;
- Individuals with diabetes;
- Individuals with lung disease.
The purpose of the survey is to let an individual know their risk relative to the general population with more detail than is available through the CDC. This is achieved by mapping questions related to an individual’s medical history into diagnosis code categories from the CCSR. We also included age and gender as well as prior year hospital inpatient or ER visits. In addition to the conditions coming from the recommendations of the CDC, we included features that our other modeling efforts surfaced as important. The mapping between the survey questions and CCSR codes is described in Table 4. To turn this into a model, we extract ICD-10 diagnosis codes from the claims in the year before the prediction date and aggregate them using the CCSR categories. We create indicator features for the presence of any code in the CCSR category.
Table 4
| Feature name | Coefficient | CCSR categories |
|---|---|---|
| Intercept | –6.74 | N/A |
| Age | 0.041 | N/A |
| Gender male | 0.171 | N/A |
| Prior admissions | 0.682 | N/A |
| Prior ER visits | 0.413 | N/A |
| COPD or emphysema, cystic fibrosis, or chronic bronchitis | 1.167 | CCSR:RSP008, CCSR:END012 |
| Asthma | 1.393 | CCSR:RSP009 |
| Obesity | 0.935 | CCSR:END009 |
| Diabetes (other than when you were pregnant) | 0.096 | CCSR:END002, CCSR:END003, CCSR:END004, CCSR:END005 |
| Hypertension (also called high blood pressure) | 0.832 | CCSR:CIR007, CCSR:CIR008 |
| Congestive heart failure | 0.982 | CCSR:CIR019 |
| Heart attack (also called myocardial infarction) | 0.159 | CCSR:CIR009, CCSR:CIR010 |
| Rheumatic heart disease | 0.788 | CCSR:CIR001, CCSR:CIR002, CCSR:CIR011, CCSR:CIR014, CCSR:CIR015 |
| Stroke | 0.285 | CCSR:CIR020, CCSR:CIR021 |
| Sickle cell anemia/HIV infection/transplant | 2.582 | CCSR:BLD005, CCSR:INF006, CCSR:FAC023 |
| Chronic kidney disease | 0.966 | CCSR:GEN003 |
| Hemodialysis | 1.369 | CCSR:GEN002 |
| Liver disease | 0.055 | CCSR:DIG019 |
| Pneumonia, acute bronchitis, influenza or other acute respiratory infection | 0.696 | CCSR:RSP002, CCSR:RSP003, CCSR:RSP005, CCSR:RSP006 |
| Cancer | 1.091 | CCSR:NEO |
| Neurocognitive conditions | 0.294 | CCSR:NVS011, CCSR:CIR022, CCSR:CIR025 |
| Pregnancy | 0.789 | CCSR:PRG |
| COPD × age | –0.002 | N/A |
| Asthma × age | –0.015 | N/A |
| Obesity × age | –0.004 | N/A |
| Diabetes × age | 0.000 | N/A |
| Hypertension × age | 0.005 | N/A |
| Congestive heart failure × age | –0.007 | N/A |
| Myocardial infarction × age | 0.003 | N/A |
| Rheumatic heart disease × age | –0.008 | N/A |
| Stroke × age | –0.003 | N/A |
| Sickle cell/HIV/Transplate × age | –0.028 | N/A |
| Chronic kidney disease × age | –0.008 | N/A |
| Hemodialysis × age | –0.018 | N/a |
| Liver disease × age | 0.001 | N/a |
| Pneumonia, acute bronchitis, influenza or other acute respiratory infection × age | –0.005 | N/a |
| Cancer × age | –0.009 | N/a |
| Neurocognitive conditions × age | 0.004 | N/a |
| Pregnancy, childbirth and the puerperium × Age | –0.003 | N/a |
The fully specified survey risk factors model. The coefficient indicates the linear regression coefficient for the features. In cases where the feature is mapped to a set of CCSR diagnosis codes, the CCSR categories column indicates which specific categories correspond to that feature. Interaction columns are listed in the form feature 1 × feature 2 and represent a multiplication of two existing features to get the feature value. The intercept column is a feature with a value that is always 1. CDC, Centers for Disease Control and Prevention; CCSR, Clinical Classifications Software Refined; ER, emergency room; COPD, chronic obstructive pulmonary disease.
A logistic regression model is then trained on the available claims data. Logistic regression was chosen specifically due to the ease with which this model can be encoded by hand. The other models we built leverage a specific software package, and their use is limited to individuals who are technically proficient with machine learning in the python programming language. By contrast, the logistic regression model can be easily implemented by hand for any individual with the ability to properly format their data. A person’s percentile risk score is based on risk relative to the other values in the training distribution. In addition to the CCSR codes, Table 4 includes the coefficients associated with these features in the logistic regression model.
Diagnosis history model
The diagnosis history model uses gradient boosted trees. Gradient boosted trees is a machine learning method that uses an ensemble of simple models to create highly accurate predictions (17). The resulting models demonstrate higher accuracy. A drawback to these models is that they are significantly more complex; consequently, “by hand” implementations of such models are impractical. A nice feature of gradient boosted trees is that they are fairly robust against learning features that are eccentricities of the training data, but do not extend well to future data. As such, we allow full diagnosis histories to be leveraged within our simpler XGBoost model. In this approach, every category in the full CCSR is converted into an indicator feature, resulting in 559 features. A 3-month delay was imposed on the claims data, so that claims within the most recent 3 months before the prediction date were not used to make the predictions. This 3-month delay simulates the delay in claims processing that usually occurs in practical settings and enables the model to be used with current claims data. This delay was not imposed in the survey risk factors model since questionnaires would generally use current data. The GitHub repository for the diagnosis history model contains scripts that automatically prepare the features for this model from simple CSV files containing claims data.
Expanded feature model
We additionally built a model within the ClosedLoop platform. The ClosedLoop platform is a software system designed to enable rapid creation of machine learning models utilizing healthcare data. The full details of the platform are outside the bounds of this paper; however, using the platform allows us to leverage multiple, complex engineered features coming from peer-reviewed studies. Examples are social determinants of health and the Charlson comorbidity index (CCI) (18). The computation of these features from claims data is often complex or involves the linking of additional data. These are operations handled by the ClosedLoop platform that are not easy to extract into a diagnosis history format, but do provide improved model accuracy. The expanded feature model uses gradient boosted trees, the same modeling method as the diagnosis history model.
Model training & cross-validation
We employ an 80-20 train-test split on the dataset. For the Healthfirst dataset, individuals contribute multiple data points. To avoid data leakage the train test split is performed at the level of an individual. This means that data points for a given person will only be incorporated into the train or test dataset. The logistic regression model was sufficiently simple that hyperparameter tuning was not performed. For the diagnosis history and expanded feature models, the model was tuned using the Bayesian hyperparameter search (19). For this search, we perform 5-fold cross validation within the test set, using the mean out of sample performance to assess model effectiveness. Once a parameter set is chosen, the full training data set is used to train the final version of the model with the optimized hyperparameters. Stated performance metrics are always with respect to the withheld test dataset.
Statistical analysis
We quantify the performance of the C-19 index using metrics that are relevant for its intended use. Specifically, we quote a ROC AUC for comparison of general models in the healthcare space. We also consider the intended use of the model, which is to identify highly vulnerable populations for additional targeting. In this case, a more appropriate metric is sensitivity at low alerts rates. This metric measures the sensitivity of the model when looking only at some small percentage of the population. To provide a baseline for comparison, the models are compared to the CCI (18). The CCI is a rules-based risk scoring algorithm based on claims data, and provides a good baseline for these types of models.
All procedures performed in this study were in accordance with the Declaration of Helsinki (as revised in 2013). Because this was a retrospective study done on anonymized data, no informed consent or ethical review was required.
Results
The standard receiver operating characteristic curves show that all three models have identical areas under the curve, at 0.87. The sensitivity at low alert rate is plotted for alert rates up to 20% of the population, and is shown in Figure 1. Additionally, the metrics quantifying the effectiveness of our models are presented in Table 5. The models outperformed the CCI, showing an increased effectiveness of this model over traditional rules-based approaches.
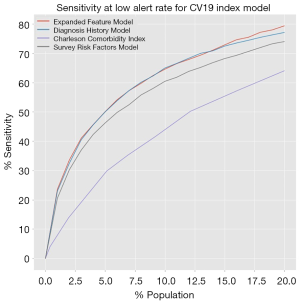
Table 5
| Model | ROC AUC | Sensitivity at 5% |
|---|---|---|
| Survey risk factors | 0.86 | 0.498 |
| Diagnosis history | 0.87 | 0.538 |
| Expanded feature | 0.87 | 0.541 |
| Charlson comorbidity | 0.79 | 0.291 |
Measures of effectiveness for the different models. The ROC AUC score is the area under the receiver-operating characteristic curve, a common accuracy statistic for binary predictors. Sensitivity at 5% indicates the percentage of condition positive examples who are identified in the top 5% of predictions.
Validation
The expanded features model has subsequently been validated by evaluating its results against approximately 14,000 hospital admissions for known COVID-19 cases in New York City from 2/1/2020 until mid-May 2020. We validated the model by comparing the mortality rate for these admissions with their predicted vulnerability. All cases were insured by Healthfirst, and each patient’s prior claims data was used to compute their predictions for the expanded features model.
Members who were admitted and survived had an average score of 2.4% and members who were admitted and passed away had an average score of 3.3% (a 38% relative difference). The ROC AUC was 0.68. This is lower than the test ROC, but that drop is expected since this test set consisted only of patients who had already been admitted for COVID-19, presumably removing many of the low vulnerability patients and increasing the difficulty of the prediction problem.
The 14,000 cases were divided into an adult Medicaid population and a Medicare population. Table 6 sorts each population by their vulnerability index and shows the death rate of each decile along with the lift relative to the overall death rate.
Table 6
| Decile | Medicaid mortality (%) | Medicaid lift | Medicare mortality (%) | Medicare lift |
|---|---|---|---|---|
| Top 10% | 6.48 | 1.30 | 5.56 | 2.99 |
| Top 20% | 6.43 | 1.29 | 3.70 | 2.00 |
| Top 30% | 6.39 | 1.29 | 3.09 | 1.66 |
| Top 40% | 6.39 | 1.28 | 2.78 | 1.50 |
| Top 50% | 6.21 | 1.25 | 2.59 | 1.40 |
| Top 60% | 5.80 | 1.17 | 3.10 | 1.67 |
| Top 70% | 5.51 | 1.11 | 2.65 | 1.43 |
| Top 80% | 5.25 | 1.06 | 2.32 | 1.25 |
| Top 90% | 5.11 | 1.03 | 2.06 | 1.11 |
| Full | 4.97 | – | 1.86 | – |
Each population in the validation set was divided into deciles based on predicted vulnerabilities. The cumulative death rate for each decile is shown in the mortality % column. The lift columns divide the death rate of the decile by the total death rate for the population. The deciles demonstrate that increased predicted COVID-19 vulnerability is in fact associated with higher rates of mortality in hospitalized patients. COVID-19, coronavirus disease 2019.
Discussion
In an effort to make these models as broadly available as possible we have provided several different avenues for the models to be used, each optimized for a different user base. The logistic regression model powers a publicly available web-based survey at http://c19survey.closedloop.ai. The diagnosis history model is available through GitHub at https://github.com/closedloop-ai/cv19index. This model is written in the Python programming language. We have included synthetic data for testing a wrapper code that converts tabular medical claims data to the input format specific for our models. We encourage the healthcare data science community to fork the repository and adapt it to their own purposes. We encourage collaboration from the open-source community, and pull requests will be considered for inclusion in the main branch of the package. Finally, for those wishing to take advantage of the expanded feature model, we are providing access to the COVID-19 model hosted on the ClosedLoop platform free of charge. Please visit https://closedloop.ai/cv19index for instructions on how to gain access.
Limitations
The approach taken in this paper has several limitations. Most notably, no actual COVID-19 cases were used in the training of the model. The usefulness of the model in predicting COVID-19 vulnerability is entirely dependent on the actual occurrence of COVID-19 matching the proxy outcome. While the logic behind these decisions is defensible and we have performed a limited validation of this proxy outcome against actual data, further evaluation is needed. As more COVID-19 case data becomes available, we expect to further validate the proxy outcome and determine if it is in fact appropriate. Eventually, enough data will be available to build models on COVID-19 vulnerability itself without having to use a proxy.
Another major limitation of these models is their reliance on claims data, which is missing much clinical detail, such as lab values. For this reason, we do not recommend using these models in inpatient settings, where more detailed clinical data is likely available. The models are most useful in a population health context where the only data available is claims data. Finally, based on medical guidance the authors have decided to exclude pediatric populations from the training and test sets for these models. At the time of development, there was so little information available on COVID-19 that we could not confidently assert that the proxy endpoint we were using was appropriate for those under 18 years of age.
Due to the early release of the first versions of these models to the open source community, several organizations were able to quickly apply the model to their populations. The authors have been in contact with several organizations that have used the models to prioritize proactive outreach towards the most vulnerable members. In many cases, these organizations had existing care management teams that were able to rapidly shift to COVID-19 by applying a different prioritization and focus to their interventions. In other cases, new phone or text messaging campaigns were developed for COVID-19 that used the C-19 index as a prioritization mechanism. As the response to the pandemic continues to develop, we will continue to update the models and provide more information on their usage.
Conclusions
This pandemic has already claimed tens of thousands of lives as of this writing, and sadly this number is sure to grow. As healthcare resources are constrained by the same scarcity constraints that affect us all, it is important to empower intervention policy with the best information possible. We have provided several implementations of COVID-19 vulnerability prediction models and means of access for those models to individuals with varying levels of technical expertise. It is our hope that by providing this tool quickly to the healthcare data science community, widespread adoption will lead to more effective intervention strategies and, ultimately, help to curtail the worst effects of this pandemic.
Acknowledgments
We would like to thank Healthfirst for their collaboration on this work and for allowing us to use insights from their data in generation of the model.
Funding: We would also like to thank Amazon Web Services for sponsoring this work with AWS platform credits.
Footnote
Data Sharing Statement: Available at http://dx.doi.org/10.21037/jmai-20-47
Peer Review File: Available at http://dx.doi.org/10.21037/jmai-20-47
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jmai-20-47). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. All procedures performed in this study were in accordance with the Declaration of Helsinki (as revised in 2013). Because this was a retrospective study done on anonymized data, no informed consent or ethical review was required.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- World Health Organization. Coronavirus disease (COVID-19) pandemic. 2020 [cited 2020 Mar 15]. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019
- Sanche S, Lin YT, Xu C, et al. High contagiousness and rapid spread of severe acute respiratory syndrome coronavirus 2. Emerg Infect Dis 2020;26:1470-7. [Crossref] [PubMed]
- Hawryluk M. Mysterious heart damage, not just lung troubles, befalling COVID-19 patients. Kaiser Health News. 2020 [cited 2020 Apr 21]. Available online: https://khn.org/news/mysterious-heart-damage-not-just-lung-troubles-befalling-covid-19-patients/
- World Health Organization. WHO Director-General’s opening remarks at the media briefing on COVID-19 - 11 March 2020. 2020 [cited 2020 Mar 15]. Available online: https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020
- Specht L. What does the coronavirus mean for the U.S. health care system? Some simple math offers alarming answers. STAT. 2020 [cited 2020 Mar 15]. Available online: https://www.statnews.com/2020/03/10/simple-math-alarming-answers-covid-19/
- Page M. Why is it so hard to calculate how many people will die from covid-19? New Scientist. 2020 [cited 2020 Mar 15]. Available online: https://www.newscientist.com/article/mg24532733-700-why-is-it-so-hard-to-calculate-how-many-people-will-die-from-covid-19/
- Zhou F, Yu T, Du R, et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 2020;395:1054-62. [Crossref] [PubMed]
- CDC. COVID-19 Response Team. Preliminary Estimates of the Prevalence of Selected Underlying Health Conditions Among Patients with Coronavirus Disease 2019 - United States, February 12-March 28, 2020. MMWR Morb Mortal Wkly Rep 2020;69:382-6. [Crossref] [PubMed]
- Wu Z, McGoogan JM. Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) outbreak in China: summary of a report of 72 314 cases from the Chinese Center for Disease Control and Prevention. JAMA 2020;323:1239-42. [Crossref] [PubMed]
- Centers for Disease Control and Prevention. Coronavirus Disease 2019 (COVID-19). 2020 [cited 2020 Feb 11]. Available online: https://www.cdc.gov/coronavirus/2019-ncov/specific-groups/high-risk-complications.html
- Hubbard RE, Peel NM, Samanta M, et al. Frailty status at admission to hospital predicts multiple adverse outcomes. Age Ageing 2017;46:801-6. [Crossref] [PubMed]
- The Healthcare Cost and Utilization Project. Clinical Classifications Software Refined (CCSR) for ICD-10-CM Diagnoses. Agency for Healthcare Research and Quality. 2020 [cited 2020 Mar 15]. Available online: https://www.hcup-us.ahrq.gov/toolssoftware/ccsr/ccs_refined.jsp
- Centers for Medicare and Medicaid Services. Limited Data Set (LDS) Files. 2017 [cited 2020 Apr 14]. Available online: https://www.cms.gov/Research-Statistics-Data-and-Systems/Files-for-Order/LimitedDataSets
- ClosedLoop.ai. COVID-19 vulnerability index. 2020 [cited 2020 Apr 14]. Available online: https://closedloop.ai/c19index/
- GitHub Inc. The COVID-19 vulnerability index (CV19 index). 2020 [cited 2020 Apr 14]. Available online: https://github.com/closedloop-ai/cv19index
- Population Health Institute. Measures & data sources. County Health Rankings & Roadmaps. University of Wisconsin, 2020 [cited 2020 Feb 1]. Available online: https://www.countyhealthrankings.org/explore-health-rankings/measures-data-sources
- James G, Witten D, Hastie T, et al. An Introduction to Statistical Learning. New York: Springer, 2013.
- Charlson ME, Pompei P, Ales KL, et al. A new method of classifying prognostic comorbidity in longitudinal studies: development and validation. J Chronic Dis 1987;40:373-83. [Crossref] [PubMed]
- Snoek J, Larochelle H, Adams RP. Practical Bayesian optimization of machine learning algorithms. In: Advances in Neural Information Processing Systems, 2012:2951-9.
Cite this article as: DeCaprio D, Gartner J, McCall CJ, Burgess T, Garcia K, Kothari S, Sayed S. Building a COVID-19 vulnerability index. J Med Artif Intell 2020;3:15.


