
Artemisia: validation of a deep learning model for automatic breast density categorization
Introduction
Breast density is the terminology used in mammography to describe the proportion between fibroglandular tissue and adipose tissue. It is estimated that 50% of women who undergo mammography examinations have dense breast patterns (1).
There is evidence that mammographic density is as strong a predictor of risk for breast cancer in African-American and Asian-American women as for white women (2). High breast density is an independent risk factor for breast cancer (3-6). Furthermore, it may link to higher percentages of interval cancers (7). Dense breast tissue can mask lesions and has a negative impact on the sensitivity of the mammography with rates ranging from 85.7% for the adipose patterns to 61% for the extremely dense patterns. It can also generate an increase in false positives from 11.2% for the non-dense patterns to 23% for dense breasts (8).
Breast density can be measured through qualitative or quantitative methods. The American College of Radiology (ACR) has established a structured system for the visual classification of breast tissue. The system applies a four letter ordinal scale to classify breast parenchyma into: (I) breast tissue almost entirely fatty, (II) scattered fibroglandular tissue, (III) heterogeneously dense parenchyma, (IV) extremely dense breast (9) (Figure 1).
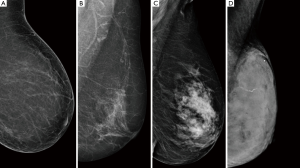
The use of ACR-BIRADS (Breast Imaging Reporting and Data System) provides a valid and reproducible application in the evaluation of breast density. However, the visual analysis is subjective, has great intra-operator and inter-operator variability and requires substantial training (10-12).
Commercial software applications such as Cumulus (13,14), LIBRA (15), Quantra (16) and Volpara (17) are based on quantitative methods for automatic breast density evaluation.
In a previous study, we had determined intra and interobserver agreement in mammographic density assessment among a group of professionals and had analyzed the agreement between experts assessment and a commercial software of a digital mammography for automatic assessment. As result, the agreement between the majority report and the commercial software was moderated (Kappa coefficient =0.43) (12).
As we mentioned before, many commercial software applications for automatic breast density evaluation are based on quantitative criteria, by this means the percentage of the breast image that is radiologically dense. Furthermore, some of the available softwares do not have disposable data regarding their validation process.
More recent machine learning methods, for example Convolutional Neural Networks (CNN), work better than traditional methods in assessing complex data like medical images (18). The application of deep learning methods for the evaluation of breast density is an area of ongoing research. Some reports about the topic have been published (19-21), but to the best of our knowledge, only Lehman’s group has applied it in daily practice (22).
Deep learning methods to assess medical images have multiple benefits; such as their reproducibility and the fact that they don’t suffer from fatigue or intraobserver variability as humans. These kinds of methods also have the ability to model the visual criteria used by professionals to categorize breast density due to their training process based on examples validated by professionals. The accuracy of diagnosis depends on the machine’s training data set, which represents the population that the machine is asked to solve. Some people may have a higher incidence of certain diseases or show different conditions. The machine must be properly trained for the target population, so it proves to be reliable in use (23).
In this article we present a CNN that had been trained based on qualitative assessment of breast density, which performance was compared to six breast images experts.
The aim of this study is to validate an algorithm of deep learning, an in-house development, for the classification of breast density according to ACR’s breast density patterns. We hypothesized that our deep learning model would achieve professional level performance.
We present the following article in accordance with the TRIPOD reporting checklist (24) (available at http://dx.doi.org/10.21037/jmai-20-43).
Methods
The study design was retrospective
We carried out this study in the Breast Diagnostic and Interventional Section of the Imaging Service in a third-level hospital. The service has been working with digital images and an integrated RIS/PACS system since 2010. The section consists of 10 specialists, 2 fellows, and reports an average of 30,000 mammograms per year. Mammographic studies are randomly assigned to medical radiologists daily for reporting. Each of them receives between 200 and 400 cases monthly. Ten percent of studies reported by specialists (approximately 300 studies monthly) and all of the studies reported by fellows are subjected to peer review. In addition, quality report audits are carried out by the doctor who requested the study.
During the validation process, we included screening mammograms performed throughout February 2019 using AMULET Innovality FUJIFilm mammogram. Each of the mammograms had an assigned breast density category by commercial software application AMULET Innovality—3000AWS7.0 Option—FUJIFilm®. From 2,640 mammograms performed throughout February, we took 451 images randomly, all from different studies and patients. We selected only mediolateral oblique or craniocaudal incidence from each study. Focused, magnified incidences and mammographic studies of patients with gigantomastia and a personal history of breast surgery (including breast implants) were excluded. Patients were between 45 and 90 years old.
Six radiologist physicians with a range of 2 to 19 years of experience participated in this study. They had to evaluate the category of mammographic density of each image. The professionals had no knowledge of the clinical history of the patients or demographic data. Besides, they were unaware of the category of breast density assigned by the other observers, including the interpretation of original medical reports as well as the evaluation of the commercial software application. For cases in which there was a tie (distribution of non-modal categorizations), a seventh imaging specialist categorized mammography to reach agreement.
Test methods
For the automatic categorization of the mammograms, we performed a convolutional neural network (CNN). In the CNN development process, we used 10,229 screening mammography images made in our hospital during the years 2017 and 2018. From those, 7,323 images were allocated to training with a homogeneous distribution among the 4 categories of breast density; 2,130 were used for the tuning phase, and 776 were used for testing. The ground truth was the category assigned in the study report. In the test phase, an accuracy of 73.32% was achieved. For development and testing, we used images from multiple manufacturers of mammographs.
The CNN architecture consists of 5 convolutional layers interspersed with 5 grouping layers, and 5 dense layers with dropout regularization. The network has 4 output nodes representing each ACR density category. The input images were preprocessed at a size of 256×256 pixels.
For development, Python 3.5 and the Keras library version 2.2.5 were used.
The CNN was baptized with the name Artemisia in honor of the Baroque painter Artemisia Gentileschi.
Once the network was developed and tested, its performance was evaluated before a group of professionals and commercial software applications.
As there was no gold standard in the evaluation of breast density, it was decided to use a reference standard that was the report of the majority of the observers. This is the statistical mode among all the categories reported by professionals (25). In the cases of a tie in the evaluation, a seventh evaluator with 7 years of experience participated to define the corresponding category. This process was repeated in two stages, with an interval of one month, in order to evaluate the variability of the majority report and the intra-observer variability (quote).
Variables of interest and statistical analysis
For each of the stages, we evaluated the matches of Artemisia, the observers, a commercial software application and the majority report, by calculating linearly weighted Kappa coefficients (k). In turn, we report the intra-observer variability for each of the participants, comparing the agreement between both stages. For the calculation we use the method described by Cohen and Fleiss (26,27) and we take as reference the subdivision of the coefficient k of Landis and Koch (0: “poor”; from 0 to 0.2: “slight”; from 0.21 to 0.4: “fair”; from 0.41 to 0.6: “moderate”; from 0.61 to 0.8: “substantial”; from 0.81 to 1: “almost perfect”) (28). Additionally, according to the clinical relevance, the results were dichotomized in non-dense pattern (A or B) and dense pattern (C or D).
The diagnostic performance of Artemisia was also evaluated in comparison with the majority report through sensitivity, specificity, positive and negative predictive value with their respective confidence intervals. We chose to report these last metrics only from the first stage. On the one hand to avoid redundancy and on the other hand because they reflect the professionals’ first encounter with the images.
For the descriptive analysis, continuous variables were reported as mean with their standard deviation. Quantitative variables were reported as a percentage and their absolute number. We report the results with a 95% confidence interval (CI). The statistical software application is STATA v. 14 and R version 3.6.0.
A sample estimate was made calculating 80% of sensitivity with a 0.05 confidence interval and a prevalence of high breast density of 50% using 451 images in total (29).
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). It was approved by the institutional ethics committee of Hospital Italiano de Buenos Aires, (NO.: CEPI #4057); and individual consent for this retrospective analysis was waived.
Results
We took the categories assigned by the six professionals, the commercial software application, the CNN and the majority report for the 451 mammographic images. The prevalence of high breast density in the sample was 41%. Table 1 of liner kappa coefficients presents the concordances between all the participants in each stage. The levels of agreement between the CNN and the majority report were k=0.64 (95% CI: 0.58–0.69) in the first stage and k=0.57 (95% CI: 0.52–0.63) in the second stage. In turn, the values of the professionals with respect to the majority report were in the ranges k=(0.64; 0.84) in the first stage and k= (0.54; 0.83) in the second stage. As for the commercial software application with respect to the majority report, the kappa for the two stages were 0.46 and 0.44, respectively. Considering the dichotomized category dense and non-dense, the agreement between the CNN and the majority report was k=0.71 (0.64–0.78) and k=0.70 (95% CI: 0.63–0.76).
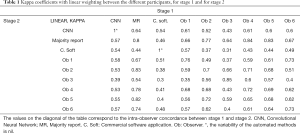
Full table
The intra-observer concordances, in other words, the coincidence between stage 1 and stage 2 for the same observer, are distributed on the diagonal of Table 1. For the six observers, the range of values for intra-observer variability is k=0.68 to 0.85. Furthermore, the majority report also shows intra-observer variability with k=0.8 (CI). In other words, the reference contains an intrinsic variability, which affects the performance evaluation in the rest of the participants. Automated methods have no intra-observer variability: the values for the CNN and the commercial software application are k=1. Because of this, the agreement between the CNN and the commercial software is the same for both stages: k=0.54 (95% CI: 0.48–0.60).
The concordances of the six professionals with the commercial software application ranges between k=0.31 and 0.57 in the first stage, and between 0.30 and 0.51 in the second stage. In which cases, for the first stage, the CNN is within the coincidence range of the professionals. In the second stage, the CNN exceeds the range of the professionals. Besides, the CNN overcomes the majority report concordance, k=0.44 and k=0.46, for each stage, respectively.
In addition, we report sensitivity, specificity, positive predictive value, negative predictive value, and the agreement of the CNN for each category, and for dichotomization in dense (C and D) and non-dense (A and B) breast tissue (Table 2). This evaluation is reported for stage 1, and with a majority report as the reference standard.
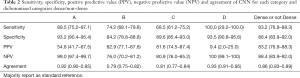

Full table
The interested reader can find the table of values with their confidence intervals in Table S1.
Discussion
Intra-observer and inter-observer variability is a well-documented problem in the evaluation of breast density proposed by ACR-BIRADS, as we have already established in our work team (12). In order to overcome this problem, we developed a convolutional neural network named Artemisia. Deep learning methods have the ability to model the visual criteria used by professionals to categorize breast density. Due to the fact that they are automated systems, they do not have intra-observer variability. We evaluated performance, analyzing the concordance between the convolutional neural network, the report of the majority, the professionals and a commercial software application. We decided to report the kappa coefficient with linear weighting, since it penalizes the disagreements between the categories proportionally to their ordinality.
The concordance between the majority report and the convolutional neural network was substantial for the first stage and moderate for the second one. Similar outcomes have been reported in different studies. Wu et al. applied a deep learning model to evaluate breast density. They reported a moderate concordance between their model and the evaluation of an experienced radiologist, with κ values of 0.48 (20). They also reported the concordances with a student (κ=0.53) and a resident (κ=0.60), demonstrating again the variability that exists in density categorization. These kappa values are not weighted. The deep learning model of Lehman group showed a substantial agreement with κ of 0.78 (95% CI: 0.73–0.82). The agreement with the original reports was κ=0.67 (22).
These studies demonstrate the feasibility of convolutional neural networks for the categorization of breast density. Deep learning algorithms achieve professional-like performances (20,30). However, regardless of the neural network architectures implemented in the different studies, the main challenge lies in mitigating variability and consequently defining and evaluating precision correctly. For this objective, we propose the comparison with the two references: the majority report and the commercial software application.
If we establish the commercial software application category as a reference standard, since it is an external invariant participant with commercial validation, we observe that the concordance of the CNN is within or above the range of professionals’ values, for each stage. This contributes to affirm that the CNN achieved a professional-like performance.
The CNN had a moderate agreement with the commercial software application. The different approaches may explain differences in categorization results. While the commercial software application applies traditional methods for processing images, our strategy is based on deep learning. Commercial software applications calculate the percentage of dense tissue and the CNN attempts to simulate the professional visual criteria. In that sense, ACR BI-RADS Atlas® 5th Edition density categories are no longer based on tissue percentage, so the visual criteria is crux (9). However, interpretability is a limiting factor of deep learning models. They also may have biases. Therefore, rigorous validation designs are necessary to mitigate these risks.
Even though ACR classifies mammographic density patterns into 4 categories, the clinical impact for patient risk management is defined by the dichotomized dense and non-dense breast patterns. We also decided to report the values in Table 2, since they are the most common metrics in the health field. When evaluating the concordance between the neural network and the majority report for dichotomous analysis of breast density between dense and non-dense, the agreement was substantial. Sensitivity and specificity have the same values as positive predictive value and negative predictive value, respectively, because the number of false positives and false negatives in the first stage are the same. Due to the very low prevalence of D pattern according to the majority report, the positive predictive value in this category is low.
As regards the sample used, it had a low number of cases with an extremely dense mammographic pattern (ACR-d), according to the prevalence reported in the institution over the last five years, during which the records were around 1–2%. Even so, the total high-density prevalence (categories c and d) in the sample was 41%, also in accordance with the prevalence in our hospital population.
The aim of these developments is to achieve an accurate categorization, as well as to reduce the variability of the labelling among professionals. There is current evidence that automated tools integrated within the workflow of medical professionals contribute to reduce variability (22). This will be our next stage. For this implementation, it will be necessary to define a categorization by study, since this validation was carried out considering the category of each image in the study, in order to compare the performance with a commercial software application.
The sample size and the randomization of the study order, avoided memory biases due to possible familiarization effects that may occur along mammography readings by the observers. Moreover, our design guaranteed the blindness of the observers to the reports of the automatic classification software and the diagnoses of the rest of the participants.
Finally, this work was carried out in a single institution. It is a reference hospital and it daily receives referrals from all over the country. A multicenter study would be convenient to evaluate this new technology.
Currently, after the validation process, Artemisia has been applied in clinical practise. Engineers had developed a solution to integrate the tool into physicians daly practise. Others studies will be needed in order to evaluate its performance in this context.
In order to access the full study protocol, please contact the corresponding author.
Conclusions
As a conclusion, considering the internal reference (majority report) and the external reference (commercial software application) and based on the concordances obtained, we can affirm that the performance of Artemisia is at the level of our professionals. These findings were what we expected, considering that Artemisia was trained with mammographies labeled by physicians of our institution.
Acknowledgments
Funding: None.
Footnote
Reporting Checklist: The authors have completed the TRIPOD reporting checklist. Available at http://dx.doi.org/10.21037/jmai-20-43
Data Sharing Statement: Available at http://dx.doi.org/10.21037/jmai-20-43
Conflict of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jmai-20-43). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). It was approved by the institutional ethics committee of Hospital Italiano de Buenos Aires (NO.: CEPI #4057); and individual consent for this retrospective analysis was waived.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Sprague BL, Gangnon RE, Burt V, et al. Prevalence of mammographically dense breasts in the United States. J Natl Cancer Inst 2014;106:dju255 [Crossref] [PubMed]
- Ursin G, Ma H, Wu AH, et al. Mammographic density and breast cancer in three ethnic groups. Cancer Epidemiol Biomarkers Prev 2003;12:332-8. [PubMed]
- Boyd NF, Martin LJ, Bronskill M, et al. Breast tissue composition and susceptibility to breast cancer. J Natl Cancer Inst 2010;102:1224-37. [Crossref] [PubMed]
- McCormack VA, dos Santos Silva I. Breast density and parenchymal patterns as markers of breast cancer risk: a meta-analysis. Cancer Epidemiol Biomarkers Prev 2006;15:1159-69. [Crossref] [PubMed]
- Boyd NF, Guo H, Martin LJ, et al. Mammographic density and the risk and detection of breast cancer. N Engl J Med 2007;356:227-36. [Crossref] [PubMed]
- Kolb TM, Lichy J, Newhouse JH. Comparison of the performance of screening mammography, physical examination, and breast US and evaluation of factors that influence them: an analysis of 27,825 patient evaluations. Radiology 2002;225:165-75. [Crossref] [PubMed]
- Holm J, Humphreys K, Li J, et al. Risk factors and tumor characteristics of interval cancers by mammographic density. J Clin Oncol 2015;33:1030-7. [Crossref] [PubMed]
- Wanders JOP, Holland K, Veldhuis WB, et al. Volumetric breast density affects performance of digital screening mammography. Breast Cancer Res Treat 2017;162:95-103. [Crossref] [PubMed]
- Sickles EA, D’Orsi CJ, Bassett LW, et al. ACR BI-RADS® Atlas, Breast imaging reporting and data system. Reston, VA: American College of Radiology 2013;39-48.
- Sprague BL, Conant EF, Onega T, et al. Variation in Mammographic Breast Density Assessments Among Radiologists in Clinical Practice: A Multicenter Observational Study. Ann Intern Med 2016;165:457-64. [Crossref] [PubMed]
- Ciatto S, Houssami N, Apruzzese A, et al. Categorizing breast mammographic density: intra- and interobserver reproducibility of BI-RADS density categories. Breast 2005;14:269-75. [Crossref] [PubMed]
- Pesce K, Tajerian M, Chico MJ, et al. Interobserver and intraobserver variability in determining breast density according to the fifth edition of the BI-RADS® Atlas. Radiologia 2020;62:481-6.
- Harvey JA, Bovbjerg VE. Quantitative assessment of mammographic breast density: relationship with breast cancer risk. Radiology 2004;230:29-41. [Crossref] [PubMed]
- Destounis S, Arieno A, Morgan R, et al. Qualitative Versus Quantitative Mammographic Breast Density Assessment: Applications for the US and Abroad. Diagnostics (Basel) 2017;7:30. [Crossref] [PubMed]
- Keller BM, Chen J, Daye D, et al. Preliminary evaluation of the publicly available Laboratory for Individualized Breast Radiodensity Assessment (LIBRA) software tool: comparison of fully automated area and volumetric density measures in a case-control study with digital mammography. Breast Cancer Res 2015;17:117. [Crossref] [PubMed]
- Richard-Davis G, Whittemore B, Disher A, et al. Evaluation of Quantra Hologic Volumetric Computerized Breast Density Software in Comparison With Manual Interpretation in a Diverse Population. Breast Cancer 2018;12:1178223418759296 [PubMed]
- Highnam R, Brady SM, Yaffe MJ, et al. Robust Breast Composition Measurement - VolparaTM. In: Digital Mammography. Berlin: Springer Berlin Heidelberg, 2010; 342-9.
- Liu Y, Chen P-HC, Krause J, et al. How to Read Articles That Use Machine Learning: Users’ Guides to the Medical Literature. JAMA 2019;322:1806-16. [Crossref] [PubMed]
- Mohamed AA, Berg WA, Peng H, et al. A deep learning method for classifying mammographic breast density categories. Med Phys 2018;45:314-21. [Crossref] [PubMed]
- Fonseca P, Castañeda B, Valenzuela R, et al. Breast Density Classification with Convolutional Neural Networks. In: Progress in Pattern Recognition, Image Analysis, Computer Vision, and Applications. Springer International Publishing, 2016; 101-8.
- Lee J, Nishikawa RM. Automated mammographic breast density estimation using a fully convolutional network. Med Phys 2018;45:1178-90. [Crossref] [PubMed]
- Lehman CD, Yala A, Schuster T, et al. Mammographic Breast Density Assessment Using Deep Learning: Clinical Implementation. Radiology 2019;290:52-8. [Crossref] [PubMed]
- Jorstad KT. Intersection of artificial intelligence and medicine: tort liability in the technological age. J Med Artif Intell 2020;3:17. [Crossref]
- Collins GS, Reitsma JB, Altman DG, et al. Transparent Reporting of a multivariable prediction model for Individual Prognosis Or Diagnosis (TRIPOD): the TRIPOD Statement. Br J Surg 2015;102:148-58. [Crossref] [PubMed]
- Ekpo EU, Ujong UP, Mello-Thoms C, et al. Assessment of Interradiologist Agreement Regarding Mammographic Breast Density Classification Using the Fifth Edition of the BI-RADS Atlas. AJR Am J Roentgenol 2016;206:1119-23.
- Cohen J. A Coefficient of Agreement for Nominal Scales. Educational and Psychological Measurement 1960;20:37-46. [Crossref]
- Fleiss JL, Cohen J. The Equivalence of Weighted Kappa and the Intraclass Correlation Coefficient as Measures of Reliability. Educ Psychol Meas 1973;33:613-9. [Crossref]
- Landis JR, Koch GG. The measurement of observer agreement for categorical data. Biometrics 1977;33:159-74. [Crossref] [PubMed]
- Jones SR, Carley S, Harrison M. An introduction to power and sample size estimation. Emerg Med J 2003;20:453-8. [Crossref] [PubMed]
- Wu N, Geras KJ, Shen Y, et al. Breast Density Classification with Deep Convolutional Neural Networks. In: 2018 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), 6682-6.
Cite this article as: Tajerian MN, Pesce K, Frangella J, Quiroga E, Boietti B, Chico MJ, Swiecicki MP, Benitez S, Rabellino M, Luna D. Artemisia: validation of a deep learning model for automatic breast density categorization. J Med Artif Intell 2021;4:5.


