
Accuracy of predicting IgHV mutation status in chronic lymphocytic leukemia using RNA expression profiling and machine learning
Introduction
The mutation status of the variable region of the immunoglobulin heavy chain (IgHV) represents one of the most widely established prognostic markers of chronic lymphocytic leukemia (CLL) (1,2). The somatic hypermutation status of the clonotypic IgHV, in particular, has been shown to underpin the risk stratification process and clinical decision-making for patients with CLL (3). The IgHV genes can be either mutated or unmutated in patients with CLL, with the latter having inferior outcomes with standard therapies (4).
This comprehension of CLL and IgHV mutation status has several clinical applications, including deducing the appropriate course of treatment for patients. An enhanced response to chemoimmunotherapy in CLL patients with mutated IgHV has been demonstrated with about 60% of patients with no evidence of disease with a plateau at 15 years (5). On the other hand, patients with unmutated IgHV have an overall inferior response to chemoimmunotherapy and a shorter time to next therapy as well as a lower overall survival (4,6,7). Jain et al. [2018] found that higher variation levels in IgHV mutations were increasingly and significantly associated with better progression-free survival and overall survival in CLL patients treated with FCR (fludarabine, cyclophosphamide and rituximab) (8). These results were replicated in the CLL8 trial, reported on by Fischer et al. [2016], which reported a significant increase in long-term remissions and overall survival in patients with mutated IgHV CLL after receiving FCR (9).
Most of the current testing for IgHV is performed using Sanger sequencing of PCR-amplified clonal or rearranged IgH variable complementarity determining regions 3 (CDR3). Sangar sequencing involves two distinct steps to determine IgHV mutation status. Firstly, clonality is detected. Second, the gene is then sequenced using Sanger sequencing and compared to predetermined germline genes obtained through immunoglobulin databases (10-12).
Despite the success and widespread acceptance of Sanger sequencing for IgHV mutation status detection, this approach of testing is not without its limitations. There are a large number of techniques available to detect IgHV mutation status; hence, discrepancies between institutions are rife.
Moreover, when using PCR, there is a risk that an alternative transcript will be amplified, giving inaccurate results. A similar disadvantage arises regarding certain primers omitting subclones (13). It has also been reported that using framework-region primers does not enable a full-length transcript to be deduced, leading to inaccuracies when calculating the percentage similarity to the homologous germline V region sequence (14). Although the availability of the immunoglobulin databases is an initial advantage when determining IgHV mutation status; this variable also poses a limitation to detection methods due to a large number of inconsistencies between the data provided. Variations may also be present in the software programs adopted to calculate the overall percentage of nucleotide mutations (13). Furthermore, it is well established that more than one clone in the CLL cell population can be seen in almost 10% of cases (15). This makes it very difficult to obtain accurate evaluation of the mutation status using Sanger sequencing.
Using NGS in sequencing can overcome most of these problems, especially when long sequence is used and covered leader region along with the other framework regions (16). NGS methodology also allows the detection of various subclones and families involved in the neoplastic process. However, NGS introduces a different set of problems, specifically determining the overall mutation status when IgHV families mutated and others that are not mutated present in the same sample. Furthermore, the IgHV mutation status sequencing does not provide any information on the presence of mutations in oncogenic genes that are relevant for evaluating the aggressiveness of the neoplastic clone.
Here we describe the use of RNA expression profiling generated from routine targeted RNA sequencing by NGS along with machine learning for the prediction of the IgHV mutation status in patients with CLL. We present the following article in accordance with the STARD reporting checklist (available at https://jmai.amegroups.com/article/view/10.21037/jmai-22-28/rc).
Methods
Study design
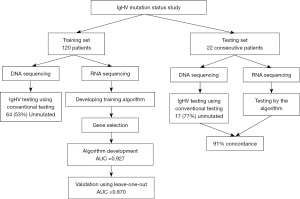
The work was retrospective study for the training set designed to develop a machine learning algorithm to explore the ability of using RNA expression profiling that is generated in the course of evaluating CLL samples in routine clinical testing to predict the IgHV mutation status. For validating this algorithm, prospective samples were tested using the developed algorithm and the conventional sequencing method (Figure 1). The goal is to eliminate the need for performing independent complicated and costly testing by sequencing IgHV locus.
Participants
Peripheral blood or bone marrow samples from a total of 142 patients with confirmed CLL by morphology and flow cytometry were studied (Figure 1). CLL diagnosis was the only required criteria for including in the study. Patients with monoclonal B-cell lymphocytosis were not included. CLL diagnosis was confirmed by morphology demonstrating small lymphocytes with scant cytoplasm and by the demonstration of the expression of CD19, CD5 and CD23 by flow cytometry. Molecular studies were also performed for evaluating various mutations especially detected in CLL (TP53, ATM, SF3B1, NOTCH1, XPO1, etc.). The samples were collected consecutively after confirming diagnosis of CLL. DNA and RNA were extracted from these samples for IgHV mutation status analysis and for RNA sequencing using a targeted 1,408 gene panel. 120 samples were used for establishing the machine learning system. This included 64 patients (53%) unmutated IgHV and 56 (47%) mutated. Twenty-two samples were used for independent blind validation of the machine learning system. This included 17 unmutated and 5 mutated. These 22 samples were collected consecutively after developing the algorithm and tested in prospective fashion. They are imbalanced, but since they are used for a secondary validation, their imbalance should not affect the algorithm. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013) and was approved by Institutional Review Board (IRB) by Western Copernicus Group (New England IRB, Aspire IRB, and Midlands IRB) (No. 1-1476184-1). Samples were collected in the course of routine clinical testing. Samples were de-identified and participants were not required to give informed consent per IRB approval.
Test methods
Determining IgHV mutation status by next generation sequencing
DNA is extracted using Maxwell RSC 48 kit (for PB/BM samples). A multiplex PCR uses primers targeting the conserved framework (FRI) and the constant region (17). All primers included are tagged with partial IDT (Integrated DNA Technology, Coralville, IA) adaptors. After first PCR, the extra-primers were removed by enzymatic digestion. The second PCR was performed using IDT adaptor/index as primers. After AMPure (Beckman Coulter, Brea, CA) wash to remove the excess primers, the quality/quantity of the library was assessed by Agilent (Santa Clara, CA) Tapestation and real time PCR. Pooling and normalization of libraries was based on real time quant. Up to 96 samples can be pooled into a one MiSeq (Illumina, San Diego, CA) run using 250×2 cycles. After MiSeq run finish, the FASTQ data were run MiXCR software, which generate IGH clonality/clonal type and quality report. All sequences from a sample obtained from MiXCR with >2% clonality were submitted to NCBI IgBlast using an automatic script (http://www.ncbi.nlm.nih/igblast/; last accessed 11/12/2021). A IgHV gene family (or families) were assigned to the patient based on the top germline IgHV gene match. A mutated status is determined when there is a >2% deviation from a germline sequence. The data generated from this testing was used as a reference standard index testing.
Targeted RNA next generation sequencing and expression profiling
The Agencourt FormaPure Total 96-Prep Kit was used to extract DNA and RNA from the same FFPE tissue lysates using an automated KingFisher Flex following the protocols recommended by the manufacturers. Samples were selectively enriched for cancer-associated genes using reagents provided in the Illumina® TruSight® RNA Pan-Cancer Panel. This panel covers 1,408 genes. cDNA was generated from the cleaved RNA fragments using random primers during the first and second strand synthesis. Sequencing adapters were ligated to the resulting double-stranded cDNA fragments. The coding regions of the expressed genes were captured from this library using sequence-specific probes to create the final library. Sequencing was performed using an Illumina NextSeq 550 system platform. Ten million reads per sample in a single run were required, and the read length was 2×150 bp. The sequencing depth was 10×–1,739×. An expression profile was generated from the sequencing coverage profile of each individual sample using Cufflinks. Expression levels were measured as fragments per kilobase of transcript per million.
Using machine learning algorithm for classifying samples
The RNA expression data of 120 samples of IgHV mutated and unmutated were used to select the proper genes that distinguish between the two groups (Figure 1). To reduce the effects of noises and avoid overfitting in selecting these genes, we employed a leave-one-out cross validation to obtain a robust performance measure. For an individual gene, a Geometric Mean Naïve Bayesian (GMNB) classifier (identical to standard Naïve Bayes for a single gene) is constructed on the training subset and tested on the testing subset. The complement of the cross-validation error rate is used as the discriminant measure for the bins.
Instead of the overall error rate, the value d takes a sum of the error rates of the individual classes. This definition would avoid the bias when the sample sizes are not balanced for different classes. The genes were ranked by d with higher values corresponding to better performing genes for classifying the two classes. To address stability issues, we used the t-test to measure the significance of a gene in separating the 2 classes. By setting a P value threshold, insignificant bins can be filtered out.
The selected genes were used to distinguish between IgHV mutated and unmutated with k-fold cross-validation procedure (with k=12). A naïve Bayesian classifier was constructed on the training of k-1 subsets and tested on the other testing subset. We applied GMNB as the classifier to predict specific class. GMNB is a generalized naïve Bayesian classifier by applying a geometric mean to the likelihood product, which would eliminate the underflow problem commonly associated with the standard naïve Bayesian classifiers with high dimensionality (18). The training and testing subsets then rotated, and the average of the classification errors was used to measure the relevancy of the gene. The classification system was trained with the selected subset of most relevant genes. The processes of Gene selection and IgHV mutation status were applied iteratively to obtain an optimal classification system and a subset of genes relevant to distinguishing between the two groups were defined and isolated.
Statistical analysis
After selecting the individual genes and specific combinations of these genes as described above using cross validation and leave one out, we used the most reliable combination of genes in developing the overall algorithm. The final model was also tested first by leave one out and then by an independent testing set. Agreement between the two testing methods was done by analyzing receiver operating characteristic (ROC) curve and the area under the curve (AUC). The above-described algorithm provided a score for the prediction of mutated CLL. The ROC curve is used to select the proper cut-off point for the proper sensitivity and specificity for distinguishing between mutated and unmutated CLL. As expected, increasing sensitivity is associated with decreasing specificity. The generated ROC curve is used to determine the proper sensitivity and specificity and the exact cut-off point for the score. The sensitivity, specificity, positive predictive value, and negative predictive value of the new test were also calculated.
Results
High specificity in the detection of unmutated IgHV in CLL
Using RNA levels, the machine learning system was trained using all the 120 cases. The training set showed a receiver operating characteristic curve with AUC of 0.927 (Figure 2A) when 23 genes were used. Using smaller or larger numbers of genes showed similar results with some variation. To validate the system and test for overfitting, we used the leave-one-out (LOO) approach. Using the 23 genes showed AUC of 0.870 (Figure 2B). Using a larger number of genes improved the prediction mildly but LOO showed significant deterioration in the prediction confirming overfitting. The 23 gene model using the scoring system generated by the algorithm, a cut-off point at 0.503 showed sensitivity of 86% (95% confidence interval: 74.5–93%) and specificity of 93% (95% confidence interval: 82–98%) (Table 1). This data indicates that 23 gene model is adequate to distinguish between mutated and unmutated. These 23 genes are listed in Table 2.
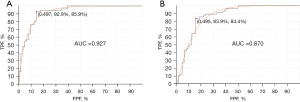
Table 1
| Detection of IgHV status | Percentage | 95% confidence interval |
|---|---|---|
| Sensitivity | 86% | 74.5–93% |
| Specificity | 93% | 82–98% |
| PPV | 93% | 83–98% |
| NPV | 85% | 73–93% |
PPV, positive predictive value; NPV, negative predictive value.
Table 2
| Order | Gene | Full name | Biological role* | Level in unmutated |
|---|---|---|---|---|
| 1 | GAB1 | Growth factor receptor bound protein 2-associated protein 1 | Member of the IRS1-like multisubstrate docking protein family. It is an important mediator of branching tubulogenesis and plays a central role in cellular growth response, transformation and apoptosis | High |
| 2 | BCL7A | B-cell CLL/lymphoma 7 protein family member A | Related to the pathogenesis of a subset of high-grade B cell non-Hodgkin lymphoma. The N-terminal segment involved in the neoplastic process via translocations | High |
| 3 | ADD3 | Adducin 3 | Membrane-cytoskeleton-associated protein that promotes the assembly of the spectrin-actin network. Plays a role in actin filament capping | Low |
| 4 | BAG4 | Bcl-2-associated athanogene 4 | An anti-apoptotic protein that functions through interactions with a variety of cell apoptosis and growth-related proteins including BCL-2, Raf-protein kinase, steroid hormone receptors, growth factor receptors and members of the heat shock protein 70 kDa family. This protein contains a BAG domain near the C-terminus, which could bind and inhibit the chaperone activity of Hsc70/Hsp70. This protein was found to be associated with the death domain of TNF-R1 and DR3, and thereby negatively regulates downstream cell death signaling | Low |
| 5 | TRAF2 | TNF receptor-associated factor 2 | A member of the TRAF protein family. TRAF proteins associate with, and mediate the signal transduction from members of the TNF receptor superfamily. This protein directly interacts with TNF receptors, and forms a heterodimeric complex with TRAF1. This protein is required for TNF-alpha-mediated activation of MAPK8/JNK and NF-κB. The protein complex formed by this protein and TRAF1 interacts with the inhibitor-of-apoptosis proteins (IAPs), and functions as a mediator of the anti-apoptotic signals from TNF receptors. The interaction of this protein with TRADD, a TNF receptor associated apoptotic signal transducer, ensures the recruitment of IAPs for the direct inhibition of caspase activation | High |
| 6 | AKT2 | AKT serine/threonine kinase 2 | A protein belonging to a subfamily of serine/threonine kinases containing SH2-like domains, which is involved in signaling pathways. The gene serves as an oncogene in the tumorigenesis of cancer cells. The encoded protein is a general protein kinase capable of phophorylating several known proteins, and has also been implicated in insulin signaling | Low |
| 7 | KLHL6 | Kelch like family member 6 | A member of the KLHL family of proteins, which is involved in B-lymphocyte antigen receptor signaling and germinal-center B-cell maturation. Naturally occurring mutations in this gene are associated with chronic lymphocytic leukemia | High |
| 8 | ZNF331 | Zinc finger protein 331 | A zinc finger protein containing a KRAB domain found in transcriptional repressors. This gene may be methylated and silenced in cancer cells | Low |
| 9 | MSI2 | Musashi-2 | An RNA-binding protein that is a member of the Musashi protein family. The encoded protein is transcriptional regulator that targets genes involved in development and cell cycle regulation | High |
| 10 | DUSP22 | Dual specificity phosphatase 22 | Enables non-membrane spanning protein tyrosine phosphatase activity and protein tyrosine kinase binding activity. Involved in several processes, including cellular response to epidermal growth factor stimulus; negative regulation of focal adhesion assembly; and negative regulation of non-membrane spanning protein tyrosine kinase activity. Acts upstream of or within negative regulation of transcription by RNA polymerase II | Low |
| 11 | APLP2 | Amyloid beta precursor like protein 2 | An APLP2, which is a member of the APP family including APP, APLP1 and APLP2. This protein is ubiquitously expressed. This protein interacts with MHC class I molecules. This protein has been implicated in the pathogenesis of Alzheimer’s disease | High |
| 12 | MAP2K1 | Mitogen-activated protein kinase kinase 1 | A member of the dual specificity protein kinase family, which acts as a MAP kinase kinase. MAP kinases, also known as ERKs, act as an integration point for multiple biochemical signals. This protein kinase lies upstream of MAP kinases and stimulates the enzymatic activity of MAP kinases upon wide variety of extra- and intracellular signals. As an essential component of MAP kinase signal transduction pathway, this kinase is involved in many cellular processes such as proliferation, differentiation, transcription regulation and development | High |
| 13 | EBF1 | EBF transcription factor 1 | Enables DNA-binding transcription activator activity, RNA polymerase II-specific and RNA polymerase II cis-regulatory region sequence-specific DNA binding activity. Predicted to be involved in positive regulation of transcription by RNA polymerase II. Predicted to act upstream of or within positive regulation of transcription, DNA-templated | Low |
| 14 | MYBL1 | MYB proto-oncogene like 1 | Enables DNA-binding transcription activator activity, RNA polymerase II-specific and RNA polymerase II cis-regulatory region sequence-specific DNA binding activity. Involved in positive regulation of transcription by RNA polymerase II | Low |
| 15 | TNFRSF10D | tumor necrosis factor receptor superfamily member 10D | A member of the TNF-receptor superfamily. This receptor contains an extracellular TRAIL-binding domain, a transmembrane domain, and a truncated cytoplasmic death domain. This receptor does not induce apoptosis, and has been shown to play an inhibitory role in TRAIL-induced cell apoptosis | High |
| 16 | RASGEF1A | RasGEF domain family member 1A | Enables guanyl-nucleotide exchange factor activity. Involved in cell migration and positive regulation of Ras protein signal transduction | High |
| 17 | PER1 | Period circadian protein homolog 1 | A member of the Period family of genes and is expressed in a circadian pattern in the suprachiasmatic nucleus, the primary circadian pacemaker in the mammalian brain. Genes in this family encode components of the circadian rhythms of locomotor activity, metabolism, and behavior. This gene is upregulated by CLOCK/ARNTL heterodimers but then represses this upregulation in a feedback loop using PER/CRY heterodimers to interact with CLOCK/ARNTL | Low |
| 18 | MAGED1 | Melanoma-associated antigen D1 | A member of the MAGE family. Although the protein encoded by this gene shares strong homology with members of the MAGE family, it is expressed in almost all normal adult tissues. This gene has been demonstrated to be involved in the p75 neurotrophin receptor mediated programmed cell death pathway | High |
| 19 | MKL1 | Megakaryoblastic leukemia 1 | Interacts with the transcription factor myocardin, a key regulator of smooth muscle cell differentiation. The encoded protein is predominantly nuclear and may help transduce signals from the cytoskeleton to the nucleus | High |
| 20 | DDX3X | DEAD-box helicase 3 X-linked | A member of the large DEAD-box protein family, that is defined by the presence of the conserved Asp-Glu-Ala-Asp (DEAD) motif, and has ATP-dependent RNA helicase activity. This protein has been reported to display a high level of RNA-independent ATPase activity, and unlike most DEAD-box helicases, the ATPase activity is thought to be stimulated by both RNA and DNA. In its nuclear roles include transcriptional regulation, mRNP assembly, pre-mRNA splicing, and mRNA export. In the cytoplasm, this protein is thought to be involved in translation, cellular signaling, and viral replication. Misregulation of this gene has been implicated in tumorigenesis | Low |
| 21 | AKT3 | AKT serine/threonine kinase 3 | A member of the AKT, also called PKB, serine/threonine protein kinase family. AKT kinases are known to be regulators of cell signaling in response to insulin and growth factors. They are involved in a wide variety of biological processes including cell proliferation, differentiation, apoptosis, tumorigenesis, as well as glycogen synthesis and glucose uptake. This kinase has been shown to be stimulated by PDGF, insulin, and IGF1 | High |
| 22 | SNX29 | Sorting nexin 29 | Enables phosphatidylinositol binding activity | High |
| 23 | FNBP1 | Formin binding protein 1 | A member of the formin-binding-protein family. Required to coordinate membrane tubulation with reorganization of the actin cytoskeleton during the late stage of clathrin-mediated endocytosis. Binds to lipids such as phosphatidylinositol 4,5-bisphosphate and phosphatidylserine and promotes membrane invagination and the formation of tubules. Also enhances actin polymerization via the recruitment of Wiskott-Aldrich Syndrome-Like/N-Wiskott-Aldrich Protein, which in turn activates the Arp2/3 complex. Actin polymerization may promote the fission of membrane tubules to form endocytic vesicles | Low |
*, data is collected from http://www.genecards.org (accessed 6/17/22). CLL, chronic lymphocytic leukemia; TNF-R1, tumor necrosis factor receptor type 1; DR3, death receptor-3; TNF, tumor necrosis factor; TRAF, TNF receptor associated factor; SH2-like, Src homology 2-like; KLHL, kelch-like; KRAB, Kruppel-associated box; APLP2, amyloid precursor- like protein 2; APP, amyloid precursor protein; MHC, major histocompatibility complex; MAP, mitogen-activated protein; ERKs, extracellular signal-regulated kinases; TRAIL, TNF-related apoptosis-inducing ligand; ARNTL, aryl hydrocarbon receptor nuclear translocator-like protein 1; CRY, cryptochrome circadian regulator; MAGE, melanoma antigen gene; PDGF, platelet-derived growth factor; IGF1, insulin-like growth factor 1; Arp2/3, actin related protein 2/3.
To validate the system, an independent set of 22 samples were first tested blindly by the algorithm and then tested by routine sequencing of the IgHV. Results were then compared between IgHV sequencing data and the algorithm data. There was 91% concordance between the two methodologies. One mutated case and one unmutated by IgHV DNA sequencing were misclassified by the RNA and machine learning approach.
Classification of CLL cases with more than one gene family
Of the 120 CLL cases, 21 (17.5%) showed the presence of more than one IgHV clone from completely different family. Additional 24 (20%) samples showed more than one clone but within the same family. Only three samples showed contradictory (mutated and unmutated) mutation status. The algorithm classified the three cases in a fashion consistent with the dominant clone (Table 3).
Table 3
| Sample | Clone fraction | IgHV family | Mutation rate, % | Algorithm classification |
|---|---|---|---|---|
| 1,554 | 57.14% | IGHV5-51 | 0.4 | Unmutated |
| 42.86% | IGHV1-18 | 5.2 | ||
| 4,455 | 38.71% | IGHV4-34 | 6.7 | Mutated |
| 3.01% | IGHV1-18 | 5.2 | ||
| 2.86% | IGHV1-8 | 0 | ||
| 2.11% | IGHV3-48 | 0 | ||
| 3,289 | 63.12% | IGHV1-8 | 0 | Unmutated |
| 36.88% | IGHV3-23 | 2 |
The algorithm classified the three cases according to the dominant clone. CLL, chronic lymphocytic leukemia; IgHV, immunoglobulin heavy chain.
Defining genes distinguishing IgHV unmutated from mutated CLL
Defining the important genes that play a major role in distinguishing IgHV unmutated CLL cases is very important not only for classification, but for understanding the biology behind the aggressive course of the unmutated CLL. The machine learning selected 23 genes that can distinguish between the mutated and the unmutated. These genes are listed in Table 2 in the order of their importance for distinguishing between the two groups. These genes are involved in the major pathways involved in cell proliferation, apoptosis, kinase signaling and migration.
Discussion
CLL is a heterogenous disease with different significant variation in its clinical course. The clinical staging system provides important prognostic information on prognosis and clinical course, but multiple additional biological and genetic markers are routinely used to guide therapeutic decisions. Deletion or mutations TP53 gene and specific cytogenetic changes are routinely used to predict prognosis and clinical course in CLL. IgHV mutational status is one of the important prognostic markers that is considered at diagnosis and for selecting therapeutic approaches. In patients that present with unmutated IgHV, novel targeted therapies should be considered as the current literature suggests these are equally effective irrespective of the patient’s IgHV status (19,20). However, therapy based on chemotherapy can be considered when CLL is mutated. These novel therapies include a vast number of treatments, such as Bruton tyrosine kinase (BTK) inhibitors, the apoptosis regulator B-cell leukemia/lymphoma 2 (BCL2) inhibitors, and phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta (PI3Kδ) inhibitors (21).
Routinely the heavy chain variable genes of the clonal B-cells are sequenced to assess the mutation status. The level of mutation defines each group. CLL with IgHV mutation ≤2% are classified as unmutated and cases with mutation rate >2% are classified as mutated (22). The presence of ≤2% divergence of the germline is accepted as a cut-off, but cases with 2-3% mutation are considered gray zone or intermediate with prognosis that is believed to be intermediate. The exception to this classification is CLL cases that express the IGHV 3–21 gene family. These cases have been shown to have clinical course similar to unmutated irrespective of the mutation status (23). Cases with IgHV 3–21 belong to stereotype subset #2, which dictates poor outcome irrespective of mutation status. Although it remains debatable, unmutated CLL are reported to be similar to pre-germinal center lymphoid cells and mutated CLL cells resemble post-germinal center cells.
Accuracy in sequencing and determining the mutation status is very important due to the clinical implication of this classification (3,4,24). However, several technical and biological issues complicate the accuracy of determining the mutation status. The presence of more than one clone or family in the clonal population makes determining the mutation status very difficult, especially when the cut-off point is so tight between 2% and 3%. NGS with its ability to delineate between various subclones and families made it easier to determine the mutation status but also introduced the problem of determining the overall mutation status when mutated clone and unmutated are detected. While the goal of testing for IgHV is to determine the clinical behavior and biology of the disease, conceptually expression profile might reflect this biology more accurately. Multiple researchers used different analysis methods (25,26), yet sequencing RNA using NGS allowed us to quantify RNA accurately and reproducibly (27). Here we used RNA levels in machine learning approach to distinguish between IgHV mutated and unmutated CLL. The approach allows us to classify CLL cases in routine molecular profiling. NGS of RNA and DNA of gene involved in the oncogenesis of CLL provides information that can be used for confirming diagnosis and predicted prognosis and response to therapy. It provides information on chromosomal structural abnormalities, mutation and now the IgHV mutation status. The use of RNA-based classification overcome the problem associated with Sanger sequencing or NGS because it is not dependent on isolating a specific clone and more reflects the biology of the disease. Furthermore, this approach overcomes the problem when mutated and unmutated clones are present in the neoplastic process.
The machine learning system selected 23 genes as best and adequate for distinguishing between mutated and unmutated CLL. Defining these genes is important because they may provide information for potential development of therapeutic approaches and targeted therapy. As shown in Table 2, these genes are involved in multiple pathways that present critical biological processes that play role in the clinical behavior of the disease. IgHV mutation status most likely reflects level of B-cell receptor (BCR) signaling status rather than oncogenic status. However, it is clear that the activation status of the BCR play a role in CLL cells survival and proliferation because therapy targeting members of this receptor pathway such as BTK is effective in treating CLL. The 23 genes selected by the algorithm to distinguish between mutated and unmutated includes genes likely involved in the BCR activation pathway (such as ADD3, TRAF2, KLHL6, etc.) and genes involved in the growth and proliferation (such as BAG4, GAB1, BCL7A and others). Thirteen of the 23 genes are overexpressed and 10 are downregulated in the unmutated CLL cases as compared with mutated CLL.
The relatively small number of cases used in establishing and confirming this model system is one of the limitations of this study. Further validation using large number of cases needs to be performed.
In summary these data provide a new method for distinguishing between mutated and unmutated CLL. This approach has multiple advantages over sequencing the IgHV and determining if mutations from germline are less or greater than 2%. However, this approach is not practical as a standalone RNA test and should be considered as a part of general overall molecular evaluation of CLL cases upon sequencing DNA and RNA to determine the various oncogenic mutations and prognostic and therapeutic RNA biomarkers. The described 23 genes should be included in such general evaluation so they can be used with the proper algorithm to evaluate the IgHV mutation status without having to perform a separate test. However, our approach should be confirmed with larger studies preferably with clinical outcome.
Acknowledgments
Funding: This work was funded by Genomic Testing Cooperative.
Footnote
Reporting Checklist: The authors have completed the STARD reporting checklist. Available at https://jmai.amegroups.com/article/view/10.21037/jmai-22-28/rc
Data Sharing Statement: Available at https://jmai.amegroups.com/article/view/10.21037/jmai-22-28/dss
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://jmai.amegroups.com/article/view/10.21037/jmai-22-28/coif). HZ, IDD, and WM are employed and own stocks at Genomic Testing Cooperative. AI receives payment for lectures from Pfizer in diffuse large B-cell lymphoma and is an advisory board member for SecuraBio, AstraZeneca, and Tg Therapeutics. MA works and owns stocks in a diagnostic company. The other authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013) and the study was approved by IRB (WCG IRB, No. 1-1476184-1). Samples were collected in the course of routine clinical testing. Samples were de-identified and participants were not required to give informed consent per IRB approval.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Davids MS, Vartanov A, Werner L, et al. Controversial fluorescence in situ hybridization cytogenetic abnormalities in chronic lymphocytic leukaemia: new insights from a large cohort. Br J Haematol 2015;170:694-703. [Crossref] [PubMed]
- Döhner H, Stilgenbauer S, Benner A, et al. Genomic aberrations and survival in chronic lymphocytic leukemia. N Engl J Med 2000;343:1910-6. [Crossref] [PubMed]
- Davi F, Langerak AW, de Septenville AL, et al. Immunoglobulin gene analysis in chronic lymphocytic leukemia in the era of next generation sequencing. Leukemia 2020;34:2545-51. [Crossref] [PubMed]
- Hamblin TJ, Davis Z, Gardiner A, et al. Unmutated Ig V(H) genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood 1999;94:1848-54. [Crossref] [PubMed]
- Gaidano G, Rossi D. The mutational landscape of chronic lymphocytic leukemia and its impact on prognosis and treatment. Hematology Am Soc Hematol Educ Program 2017;2017:329-37. [Crossref] [PubMed]
- Damle RN, Wasil T, Fais F, et al. Ig V gene mutation status and CD38 expression as novel prognostic indicators in chronic lymphocytic leukemia. Blood 1999;94:1840-7. [Crossref] [PubMed]
- Lin KI, Tam CS, Keating MJ, et al. Relevance of the immunoglobulin VH somatic mutation status in patients with chronic lymphocytic leukemia treated with fludarabine, cyclophosphamide, and rituximab (FCR) or related chemoimmunotherapy regimens. Blood 2009;113:3168-71. [Crossref] [PubMed]
- Jain P, Nogueras González GM, Kanagal-Shamanna R, et al. The absolute percent deviation of IGHV mutation rather than a 98% cut-off predicts survival of chronic lymphocytic leukaemia patients treated with fludarabine, cyclophosphamide and rituximab. Br J Haematol 2018;180:33-40. [Crossref] [PubMed]
- Fischer K, Bahlo J, Fink AM, et al. Long-term remissions after FCR chemoimmunotherapy in previously untreated patients with CLL: updated results of the CLL8 trial. Blood 2016;127:208-15. [Crossref] [PubMed]
- Ghia P, Stamatopoulos K, Belessi C, et al. ERIC recommendations on IGHV gene mutational status analysis in chronic lymphocytic leukemia. Leukemia 2007;21:1-3. [Crossref] [PubMed]
- Szankasi P, Bahler DW. Clinical laboratory analysis of immunoglobulin heavy chain variable region genes for chronic lymphocytic leukemia prognosis. J Mol Diagn 2010;12:244-9. [Crossref] [PubMed]
- Heather JM, Chain B. The sequence of sequencers: The history of sequencing DNA. Genomics 2016;107:1-8. [Crossref] [PubMed]
- Vollbrecht C, Mairinger FD, Koitzsch U, et al. Comprehensive Analysis of Disease-Related Genes in Chronic Lymphocytic Leukemia by Multiplex PCR-Based Next Generation Sequencing. PLoS One 2015;10:e0129544. [Crossref] [PubMed]
- Crombie J, Davids MS. IGHV mutational status testing in chronic lymphocytic leukemia. Am J Hematol 2017;92:1393-7. [Crossref] [PubMed]
- Langerak AW, Davi F, Ghia P, et al. Immunoglobulin sequence analysis and prognostication in CLL: guidelines from the ERIC review board for reliable interpretation of problematic cases. Leukemia 2011;25:979-84. [Crossref] [PubMed]
- Scheijen B, Meijers RWJ, Rijntjes J, et al. Next-generation sequencing of immunoglobulin gene rearrangements for clonality assessment: a technical feasibility study by EuroClonality-NGS. Leukemia 2019;33:2227-40. [Crossref] [PubMed]
- van Dongen JJ, Langerak AW, Brüggemann M, et al. Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 2003;17:2257-317. [Crossref] [PubMed]
- Albitar M, Zhang H, Goy A, et al. Determining clinical course of diffuse large B-cell lymphoma using targeted transcriptome and machine learning algorithms. Blood Cancer J 2022;12:25. [Crossref] [PubMed]
- Skånland SS, Mato AR. Overcoming resistance to targeted therapies in chronic lymphocytic leukemia. Blood Adv 2021;5:334-43. [Crossref] [PubMed]
- Brander D, Islam P, Barrientos JC. Tailored Treatment Strategies for Chronic Lymphocytic Leukemia in a Rapidly Changing Era. Am Soc Clin Oncol Educ Book 2019;39:487-98. [Crossref] [PubMed]
- Patel K, Pagel JM. Current and future treatment strategies in chronic lymphocytic leukemia. J Hematol Oncol 2021;14:69. [Crossref] [PubMed]
- Chiorazzi N. Implications of new prognostic markers in chronic lymphocytic leukemia. Hematology Am Soc Hematol Educ Program 2012;2012:76-87. [Crossref] [PubMed]
- Baliakas P, Agathangelidis A, Hadzidimitriou A, et al. Not all IGHV3-21 chronic lymphocytic leukemias are equal: prognostic considerations. Blood 2015;125:856-9. [Crossref] [PubMed]
- Gupta SK, Viswanatha DS, Patel KP. Evaluation of Somatic Hypermutation Status in Chronic Lymphocytic Leukemia (CLL) in the Era of Next Generation Sequencing. Front Cell Dev Biol 2020;8:357. [Crossref] [PubMed]
- Yepes S, Torres MM, Andrade RE. Clustering of Expression Data in Chronic Lymphocytic Leukemia Reveals New Molecular Subdivisions. PLoS One 2015;10:e0137132. [Crossref] [PubMed]
- Friedman DR, Weinberg JB, Barry WT, et al. A genomic approach to improve prognosis and predict therapeutic response in chronic lymphocytic leukemia. Clin Cancer Res 2009;15:6947-55. [Crossref] [PubMed]
- Mosquera Orgueira A, Antelo Rodríguez B, Alonso Vence N, et al. Time to Treatment Prediction in Chronic Lymphocytic Leukemia Based on New Transcriptional Patterns. Front Oncol 2019;9:79. [Crossref] [PubMed]
Cite this article as: Charifa A, Zhang H, Pecora A, Ip A, De Dios I, Ma W, Leslie LA, Feldman T, Goy A, Albitar M. Accuracy of predicting IgHV mutation status in chronic lymphocytic leukemia using RNA expression profiling and machine learning. J Med Artif Intell 2022;5:10.


